Research
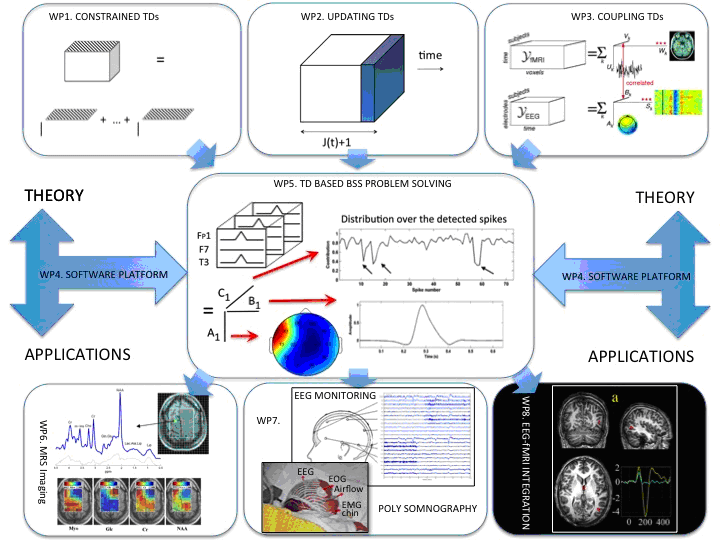
The research program is subdivided in 8 Work Packages (see Figure).
Work Packages 1 to 4 are algorithm and software oriented and aim at extending current knowledge w.r.t. tensor methods. Lieven De Lathauwer, colleague and main collaborator of the PI, will contribute as expert. WP4 serves as a common easy-to-use Matlab-based software platform integrating all relevant results.
Work Packages 5 to 8 are more application oriented. WP5 bridges the gap between theory and applications by showing how to translate core problems in biomedical signal processing into tensor based BSS problems. These formulations and corresponding algorithms are then validated for three applications
- WP1: Computing (constrained) tensor decompositions
- WP2: Updating tensor decompositions
- WP3: Coupling tensor decompositions
- WP4: Software platform for tensor based biomedical blind source separation
- WP5: Tensor based BSS problem formulations in biomedical signal processing
- WP6: Tensor based BSS in Magnetic Resonance Spectroscopic Imaging
- WP7: Tensor based BSS in functional monitoring
- WP8: Tensor based BSS in simultaneous EEG-fMRI integration
An introductory presentation to the BIOTENSORS project can be found here.
Summary of the research work done till mid-term (April 1, 2014 – April 1, 2019)
The quest for a general functional tensor framework for blind source separation
Our overall objective is the development of a general functional framework for solving tensor based blind source separation (BSS) problems in biomedical data fusion, using tensor decompositions (TDs) as basic core. We claim that TDs will allow the extraction of fairly complicated sources of biomedical activity from fairly complicated sets of uni- and multimodal data.
Significant progress is made on the development of algebraically and numerically well-founded tensor techniques for BSS, including the Canonical Polyadic Decomposition (CPD) and Block Term Decomposition (BTD) and their extensions involving block terms, various constraints, source models and heterogeneous coupled data sets. In particular:
- Strongly relaxed uniqueness conditions, both algebraic and generic, have been derived for several tensor decompositions, including block terms, constraints and coupling.
- New signal separation techniques and corresponding structured TDs have been developed, by modeling sources as exponential polynomials, rational functions and Kronecker-products. The latter yielded a novel segmentation strategy, enabling large-scale BSS and convolutive blind source identification. The structured tensor framework elegantly exploits structure in these tensors, enabling constrained decompositions of large-scale compressible tensors.
- Combination of stochastic gradient descent and tensor methods leads to a randomized block sampling algorithm allowing gigabyte and terabyte tensors to be decomposed on a laptop. Additional developments in incomplete tensor decompositions with possible linear constraints allow tensors with O(1018) entries to be compressed using only 105 samples and 4500 variables. Using polynomial constraints, high accuracy models can be reconstructed using few samples.
- A new multilinear classification paradigm has been presented based on linear systems of which the solution has a CPD structure. This has lead to new applications in, e.g., face recognition, irregular heartbeat classification and blind system identification.
- A nonlinear Least-Squares algorithm for CPD updating in one or more tensor modes enables real-time tracking of varying tensor decompositions. By only storing the new slice and the previous model (instead of the data), large-scale tensors can be decomposed using a low memory footprint.
Tensorlab efficiently implements the above algorithms. Its user friendliness has improved significantly by simplifying model construction and adding demos and Matlab-based GUIs for CPD and data compression use and tensor-based biomedical BSS. These improvements greatly facilitate non-expert usage and allow to face current grand challenges in biomedical data fusion. The following outcomes are mentioned:
- Using structured CPD/BTD, novel tensor-based classifiers for mobile Brain-Computer Interfacing (BCI) have been developed without subject-specific calibration phase: the latter showing a major benefit over traditional supervised methods at comparable accuracy.
- The added value of coupled matrix-tensor factorisations (CMTF) is shown in epileptic EEG-fMRI: the sources corresponding to the onset (spike) and propagation (slow wave) can be separated, enabling to better localize the seizure onset zone. By adding a Toeplitz factor, CMTFs allow to perform both BSS (estimation of spatial-temporal-spectral activations) but also blind system identification (estimation of neurovascular coupling or transfer function between modalities).
- Löwner tensor modeling of Magnetic Resonance (MR) Spectroscopic Imaging (MRSI) signals is shown to suppress residual water all-at-once, while non-negative CPD algorithms enable tissue type differentiation in brain tumors and detection of Multiple Sclerosis lesions, using multiparametric MRI, thereby outperforming the matrix-based counterparts.
- Various tensor-based approaches, all efficiently implemented in Tensorlab, have been applied successfully in diverse biomedical BSS problems in ECG, EEG, and MRSI signal processing. The added value of BTD versus CPD to detect seizure patterns evolving in space or frequency was proven. In addition, the power of tensor-based approaches in abnormal heartbeat detection using ECG has been shown.